
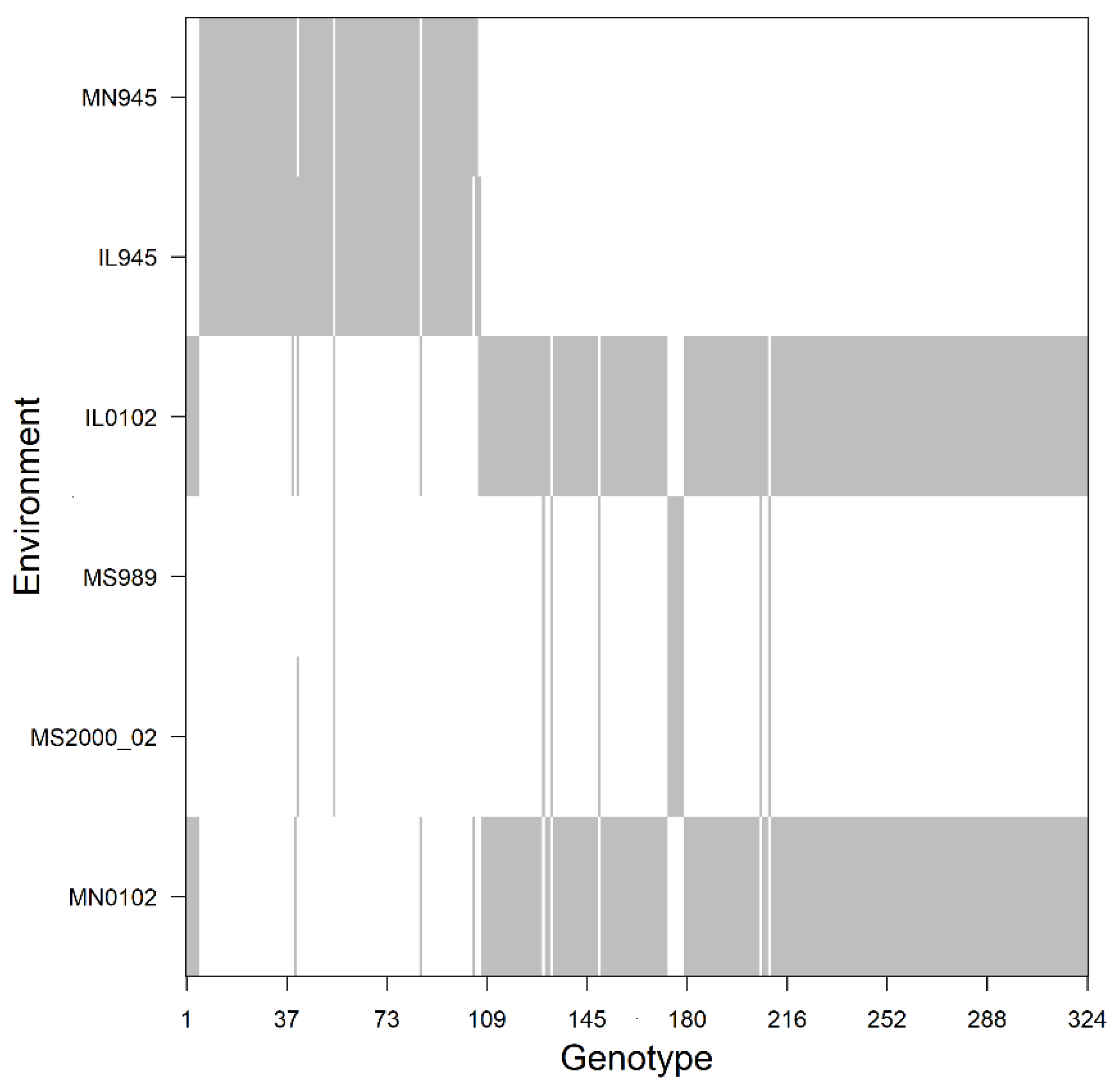
This indicates the power of the SPA model in dealing with spatial trends. In addition, the heritability, selective accuracy, and selection gain were higher when the SPA models were used. The likelihood ratio test showed that some effects changed regarding significance when the SPA and NSPA models were used. In the joint analysis, the compound symmetry structure for the genotypic effects presented the best fit. Based on the Bayesian information criteria, the SPA models were used to analyze trials E3 and E4, while the NSPA model was used for analyzing trials E1 and E2. Then, the rows and columns factors were included in the fixed and random parts of the model. The SPA models accounted for autocorrelation among rows and columns by the inclusion of first-order autoregressive matrices (AR1 ⊗ AR1). The trials consisted of 78 inter-populational maize hybrids, tested in four environments (E1, E2, E3, and E4), with three replications, under a randomized complete block design.
#Asreml r 3 manual trial
The objective of this study was to compare the spatial (SPA) and non-spatial (NSPA) models in diallel multi-environment trial analyses in maize breeding. Spatial analyses can correct spatial trends, which allow for an increase in selective accuracy. The first few lines of the phenotypic dataset $\\texttt$), and their precision will depend on the amount of information a parent has (_i.e.Spatial trends represent an obstacle to genetic evaluation in maize breeding. We also have pedigree information for the 43 parents. In this dataset, each family is formed by between 1 and 16 individuals (with an average of 12.2). There were no reciprocal or self-pollinated crosses planned, but these can occur in other crops and they might need to be identified and modelled properly. In this dataset we have a total of 71 families that originated from 43 parents however, 20 of those parents were used as both males and females in different crosses. We will be using the adjusted mean values for this trait.
#Asreml r 3 manual full
A subset of the full dataset, corresponding to diameter at breast height (DBH, inches) measured at 6 years since planting at the Nassau (Florida, USA) site, is used here.
#Asreml r 3 manual series
Individuals from these families were vegetatively propagated (cloned) and established in a series of field trials. Parents were crossed in a circular mating design, constituting several full-sib families. The data used here originates from a loblolly pine clonal study published by Resende _et al._ (2012). We will illustrate this here using an example of loblolly pine (_Pinus taeda_) in which some individuals, depending on the availability of pollen or flowers, were used in several artificial crosses as both male and female in the breeding program. This is done by _overlaying_ design matrices of the factors associated with male and female parents.

In quantitative genetic analyses, monoecious species present a particular challenge, as a given parent can contribute to the estimation of its breeding value (or GCA) as both a male and a female, something that needs to be taken into consideration when a statistical model is fitted. Some examples of monoecious species are corn, squash, banana, and many conifers, particularly those of the genus _Pinus_. In contrast, dioecious species have distinctive male and female plants. However, several commercial plant species are monoecious, which means that a given genotype will bear both male and female flowers. In most cases it is easy to assign the sex of a given individual. In many plant breeding programs, a parent is considered in several crosses. These BLUPs, which are the _general combining ability_ (GCA), or 1/2 of the _breeding value_ (BV, with BV = 2 ×× GCA) of each parent, are then used to select the best parents for future crosses or operational deployment. The progeny are later evaluated in a field experiment, and this information is used to assess the genetic worth of the parents by fitting parental linear mixed models (LMMs) and obtaining best linear unbiased predictions (BLUPs).

Most breeding programs plan several controlled crosses between outstanding parents to detect favorable alleles in their offspring.
